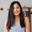Posters
Chat to the author either in person, on Gather, or on Slack during poster sessions and coffee breaks.
Join Online
#1Building the Flomics bioinformatics infrastructure from the ground up
Ecosystem

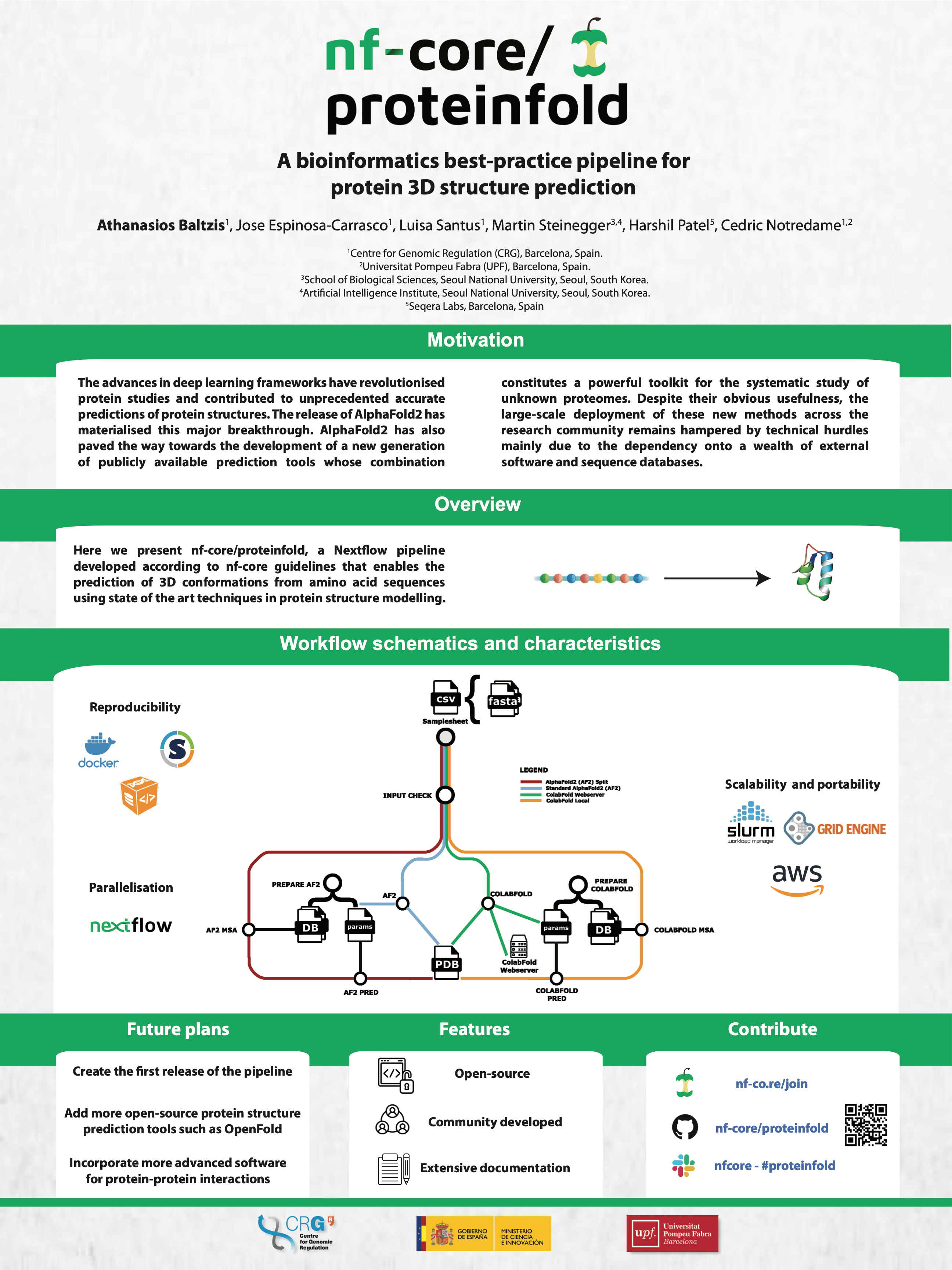
#2nf-core/proteinfold: a bioinformatics best-practice pipeline for protein 3D structure prediction
Community

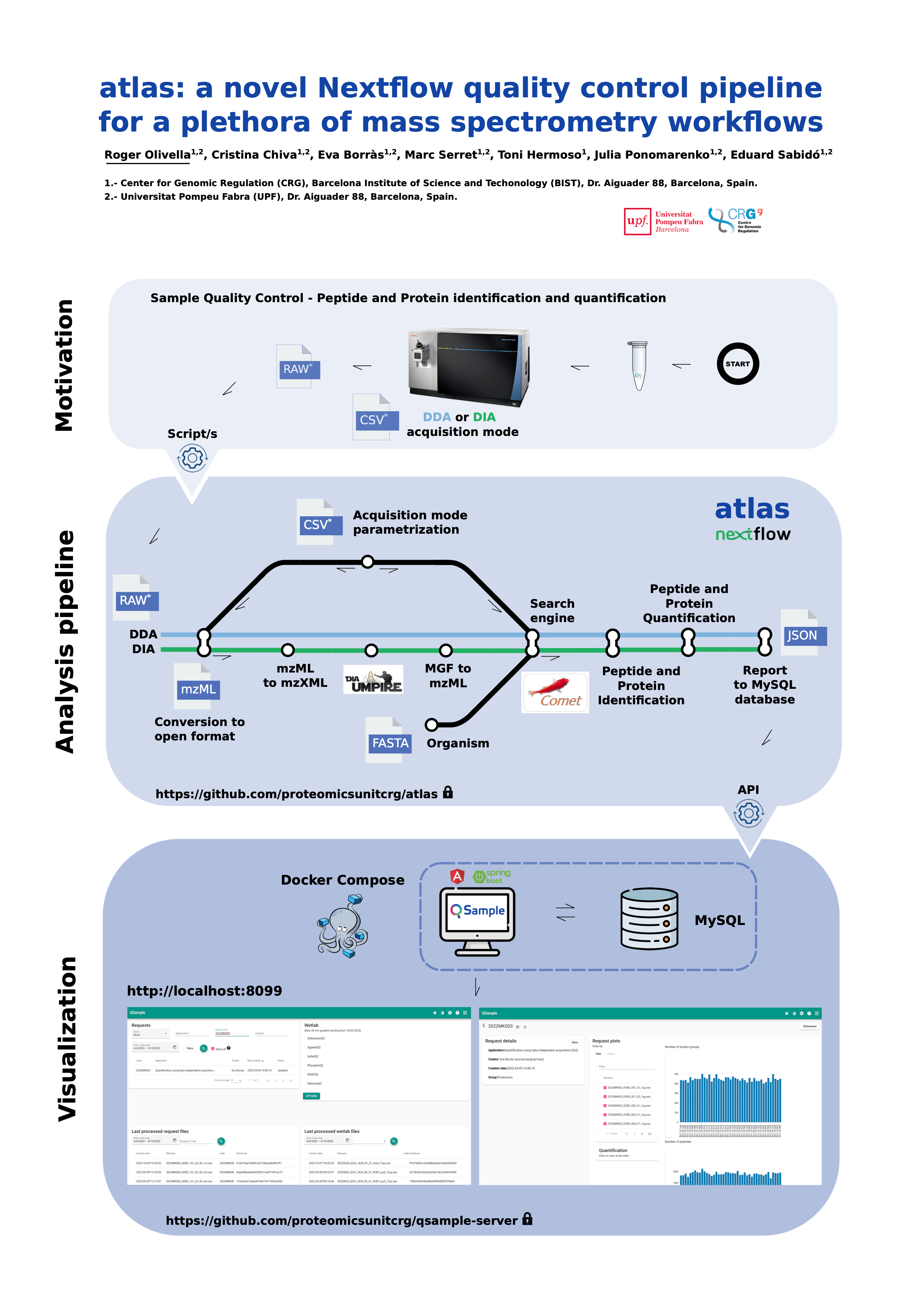
#3atlas: a novel Nextflow quality control pipeline for a plethora of mass spectrometry workflows
Community

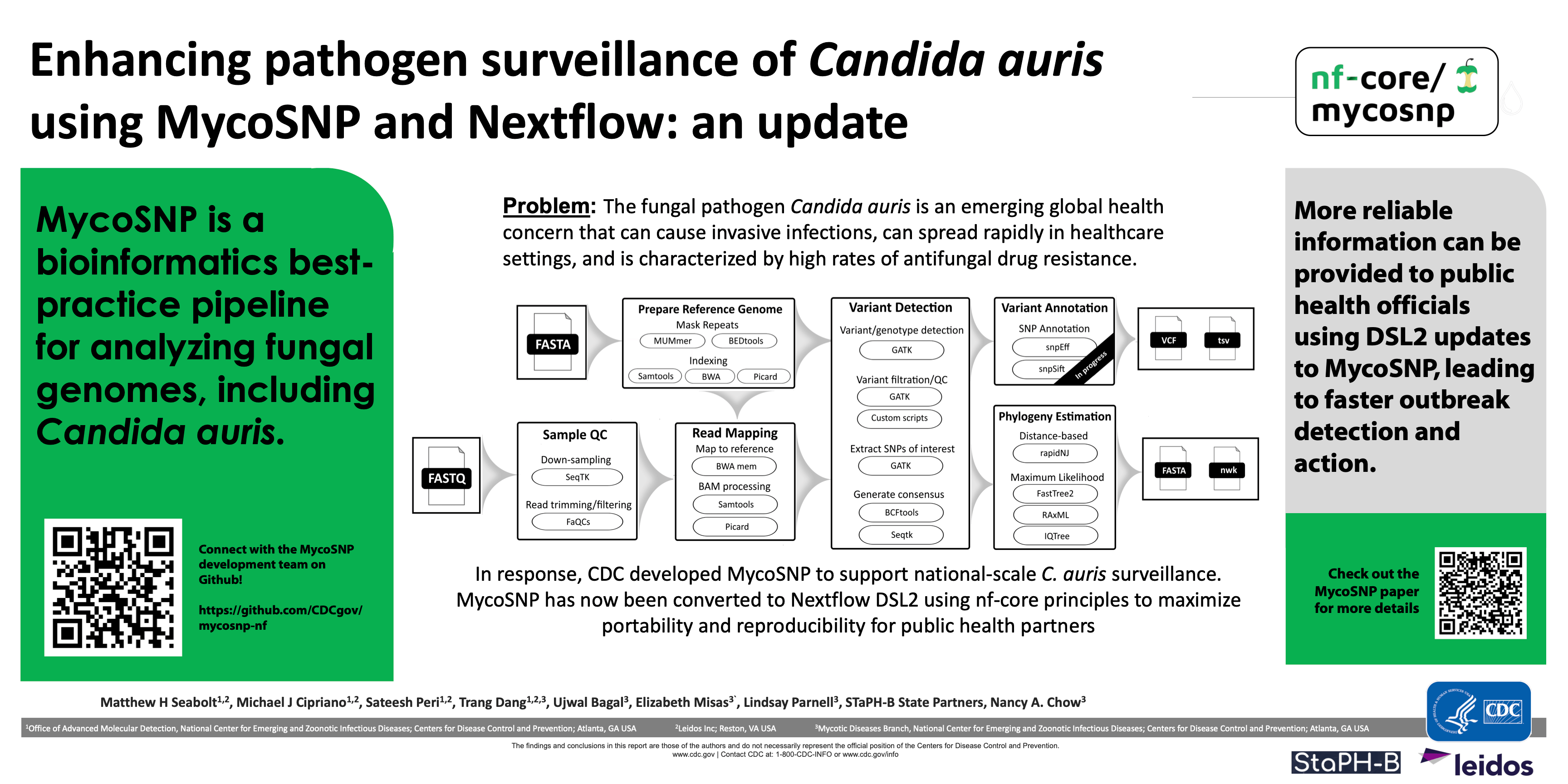
#4Enhancing pathogen surveillance of Candida auris using MycoSNP and Nextflow: an update
Community

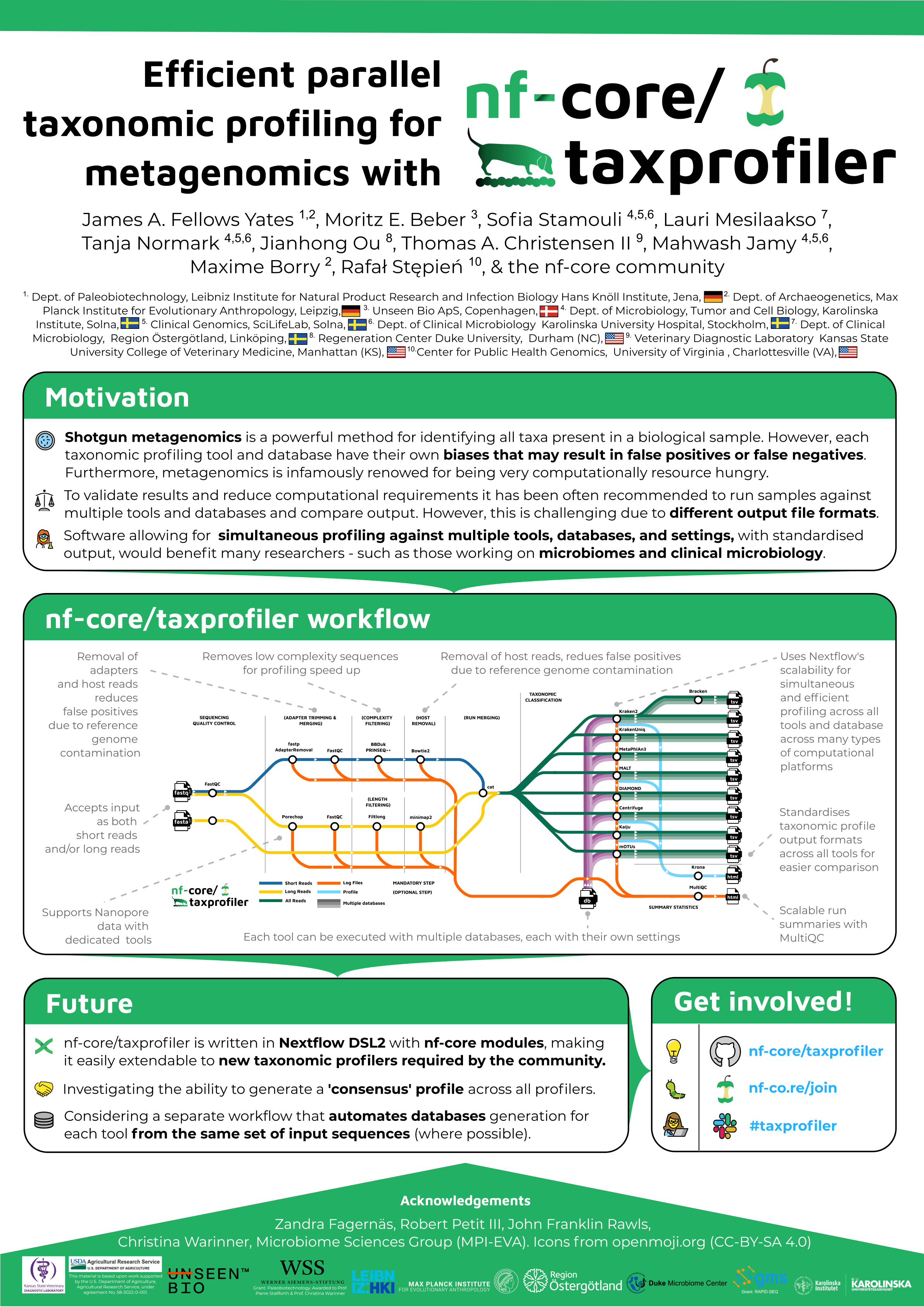
#6Efficient parallel taxonomic profiling for metagenomics with nf-core/taxprofiler
Community
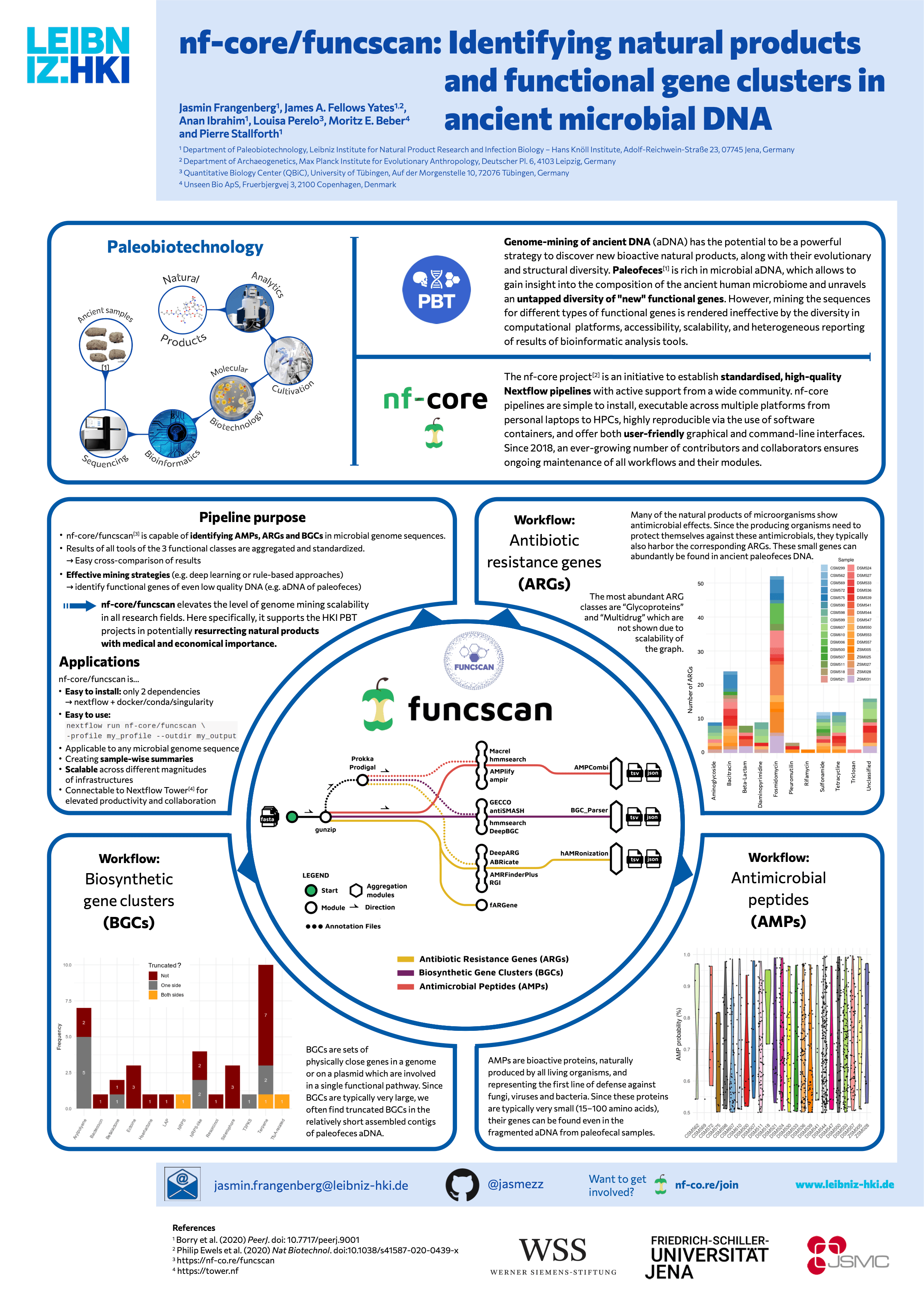


#7nf-core/funcscan: Identifying natural products and functional gene clusters in ancient microbial DNA
Community

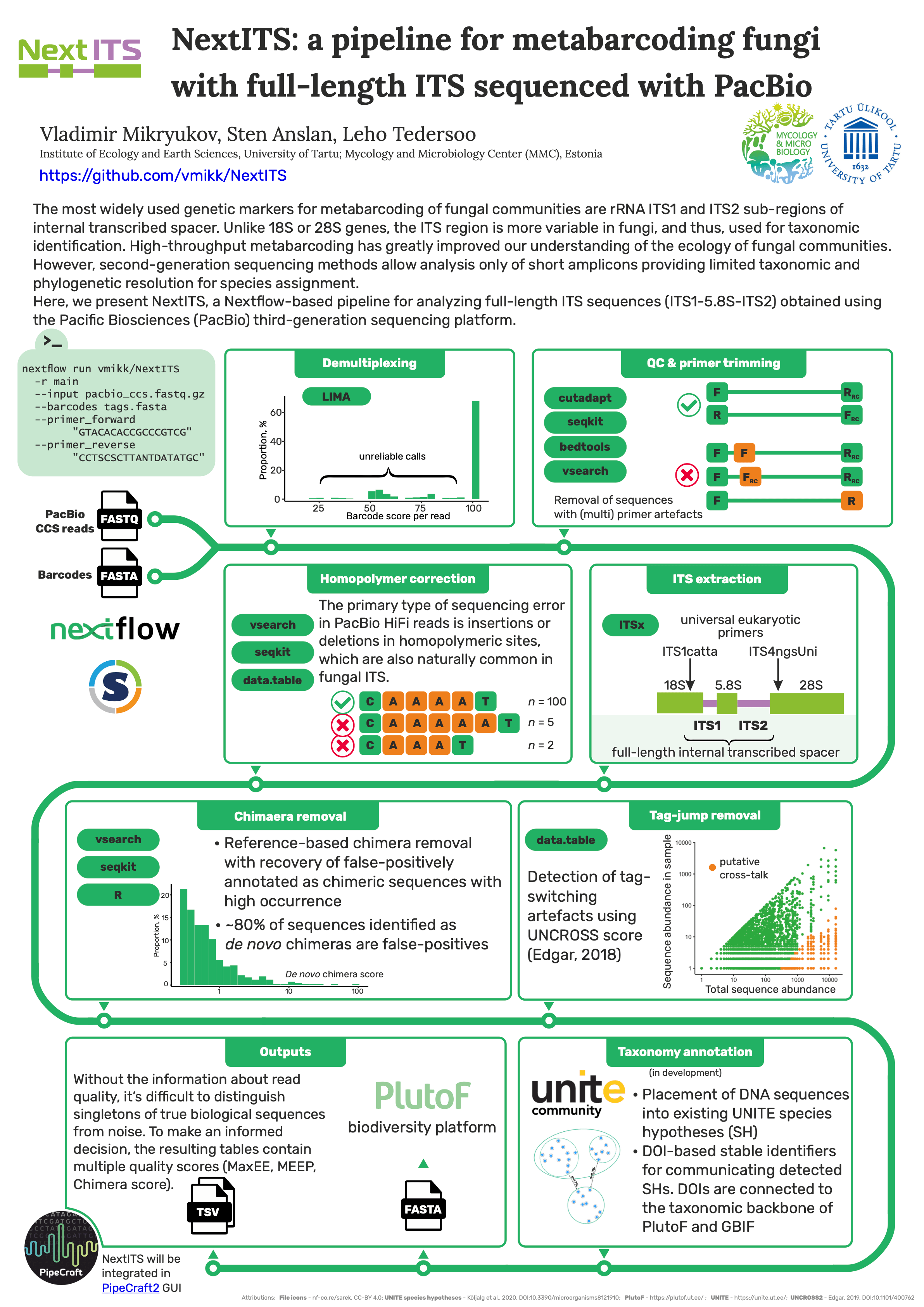
#12NextITS: a pipeline for metabarcoding fungi with full-length ITS sequenced with PacBio
Community


#13PhyloNext: a pipeline for phylogenetic diversity analysis of GBIF-mediated data in the cloud
Community
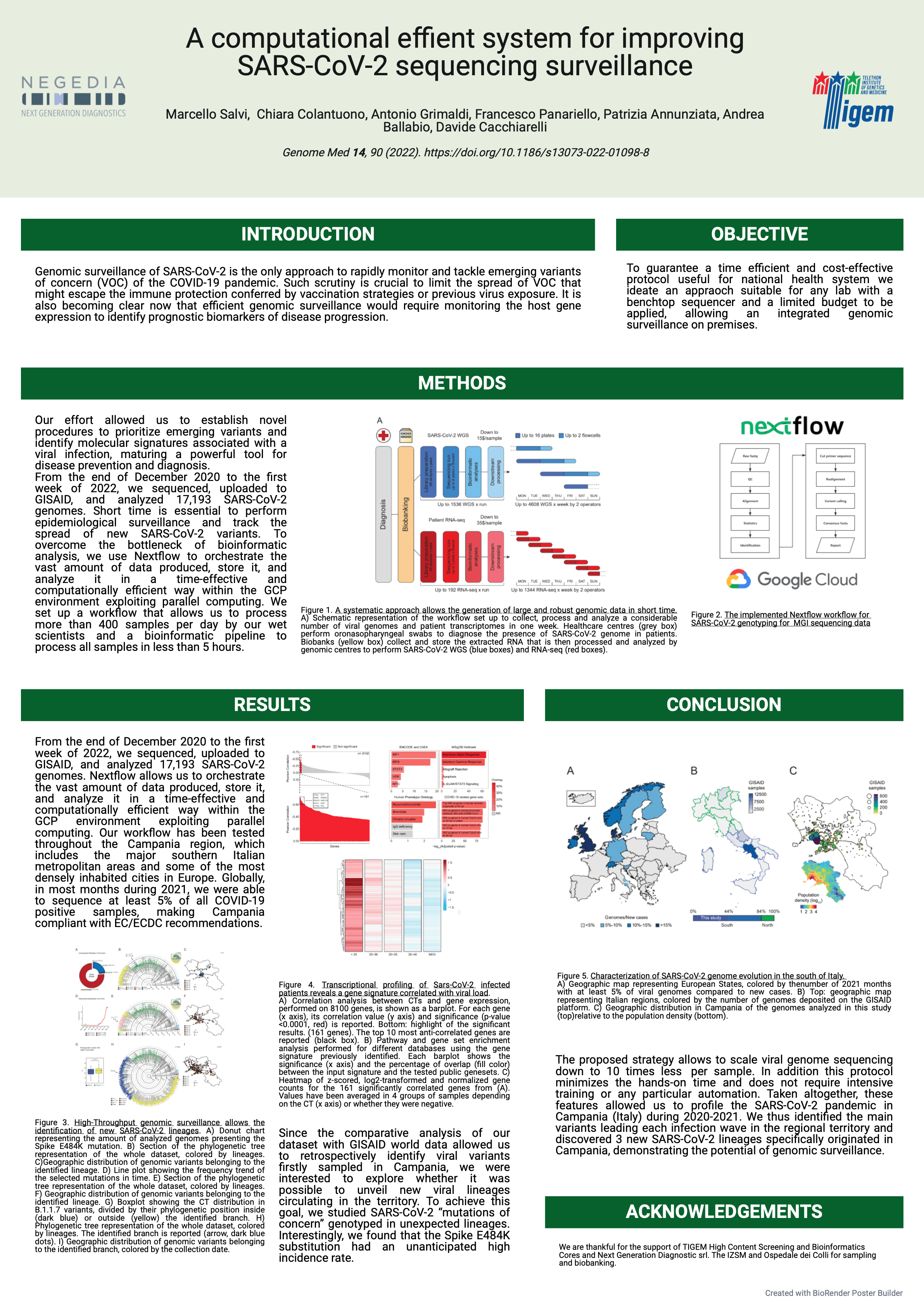


#15A computationally efficient system for improving SARS-CoV-2 sequencing surveillance
Community

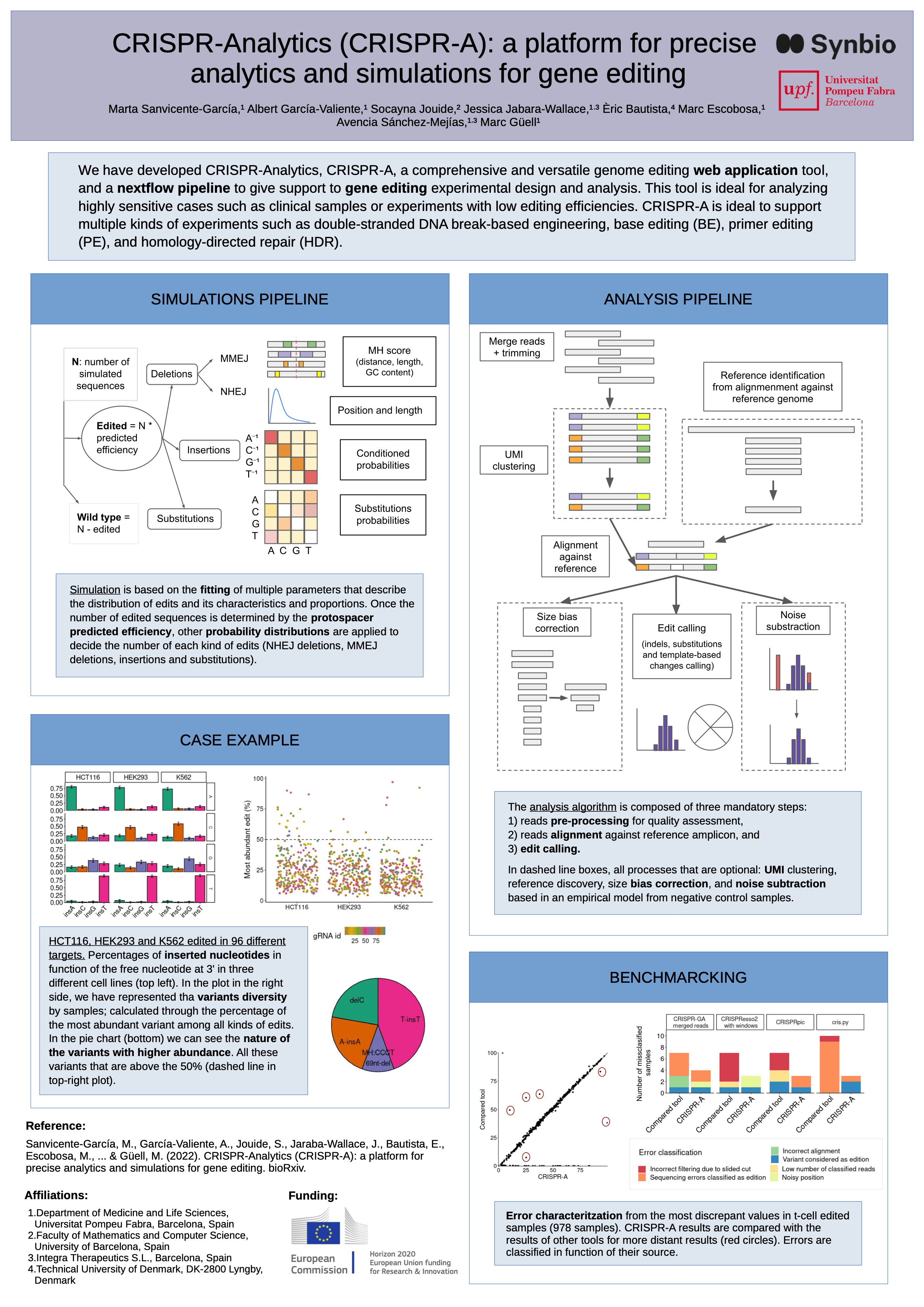
#17CRISPR-Analytics (CRISPR-A): a platform for precise analytics and simulations for gene editing
Community


#18Leveraging Nextflow and Bactopia for automated plant pathogen characterization
Community

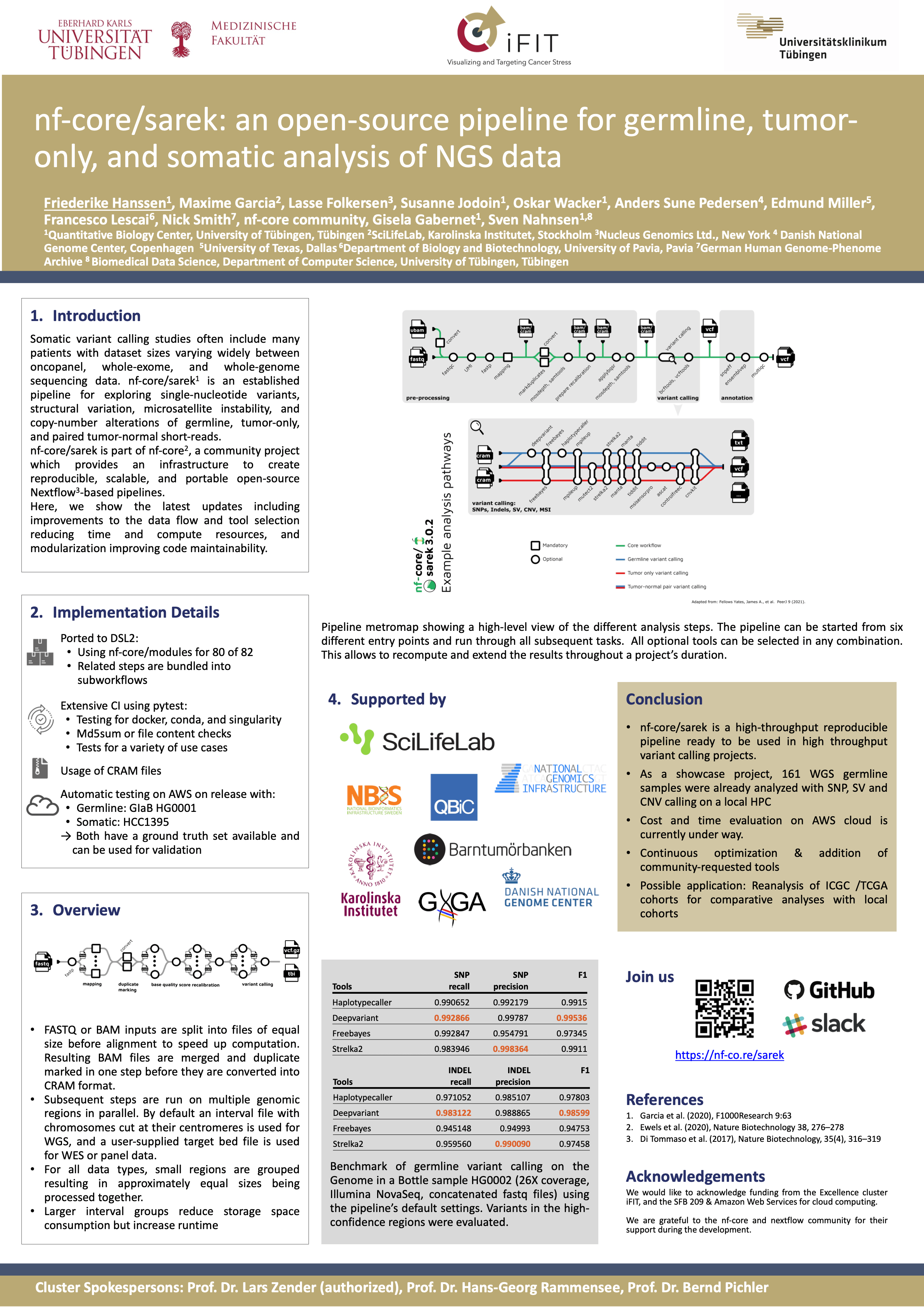
#20nf-core/sarek: A workflow for germline, tumor-only, and somatic analysis of NGS data
Community

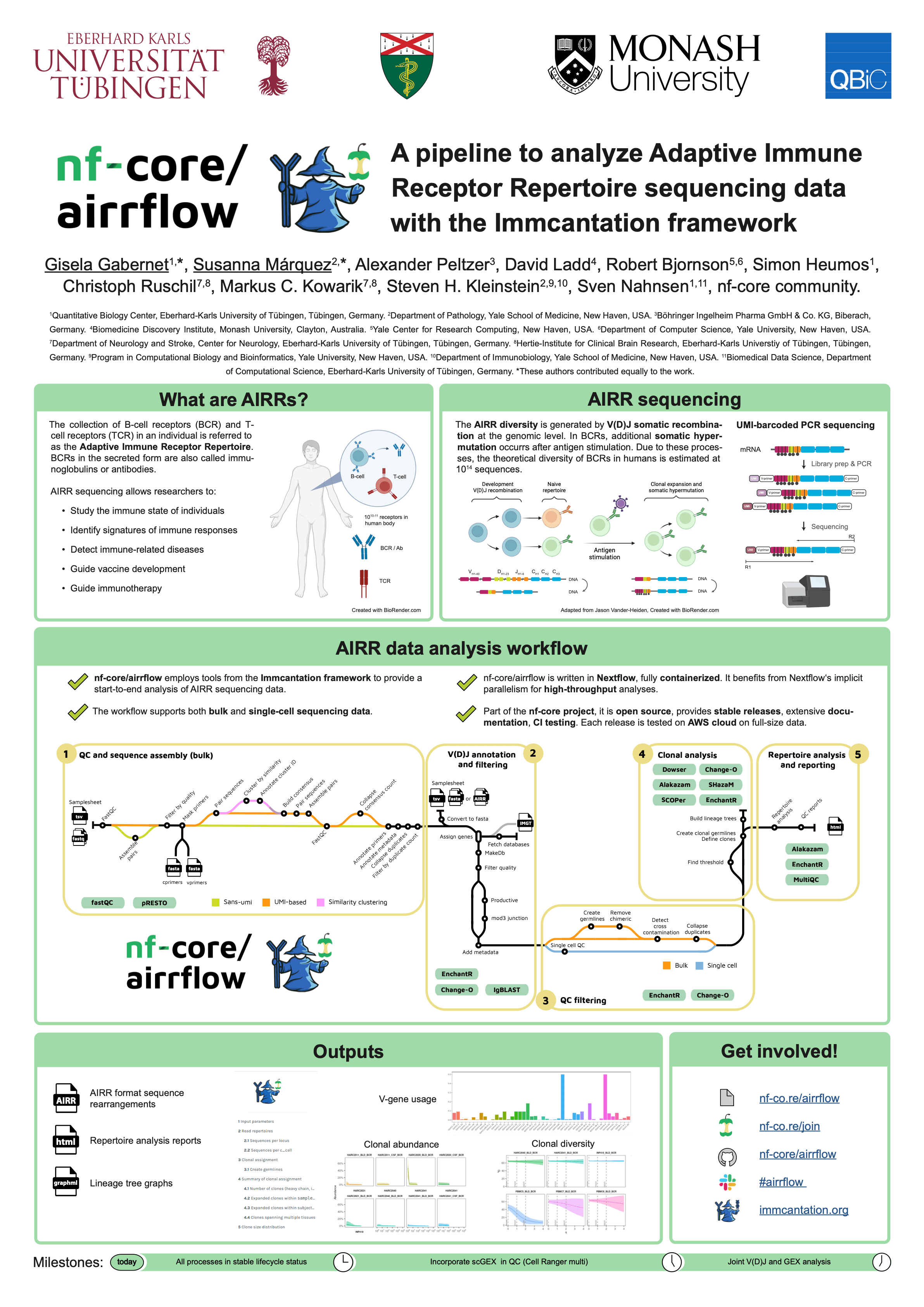
#21nf-core/airrflow: A pipeline to analyze Adaptive Immune Receptor Repertoires (AIRRs)
Community